
A small stretch of DNA that evolved in humans may help explain why our brains became so large and so powerful.
The DNA segment acts like a volume knob, dialing up production of brain cells during early development and helping build the thick, folded cortex that supports human thought, language, and reasoning.
The findings were published March 19 in Cell Stem Cell by a Duke University School of Medicine team led by developmental neurobiologist Debra Silver, PhD, and graduate student Federica Mosti, PhD. The researchers traced how a human-specific DNA sequence changes gene activity at a critical stage of brain development.

The sequence, known as HAR1984, belongs to a group called human accelerated regions, or HARs. These stretches of DNA remained nearly unchanged across mammals for millions of years, then evolved rapidly in humans.
Scientists have long known that the human brain is unusually large for our body size, with much of that difference emerging during early development, said Silver. What’s been missing, she said, is a clear explanation of how human-specific DNA changes translate into differences in brain structure.
HAR1984 appears to be a key peice, said Mosti, the first study author. Researchers have identified thousands of DNA sequences found only in humans and are only beginning to see how they shape brain development.
Scientists zero in on one switch
HARs can act like genetic switches but proving how they do so has been difficult.
To study HAR1984’s role in early cortical development, the team reached for science’s sharpest instruments — CRISPR editing and lab-grown brain organoids, work that became the center of Mosti’s graduate studies and thesis.
They swapped the human version of HAR1984 into chimpanzee organoids, and the chimp version into the human ones.
The results were striking. Chimp organoids carrying the human DNA produced more neurons and more neural progenitors, the dividing cells that generate neurons and glia to give the brain structure. Human organoids carrying the chimp version produced fewer.
The same pattern showed up in mice. Animals engineered with the human version of HAR1984 developed thicker cortices and even small folds, an unusual feature because mouse brains are typically smooth.
Bringing distant genes into close conversation
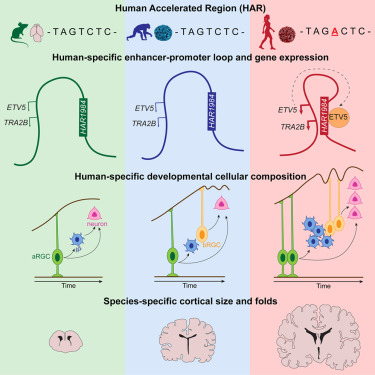
HAR1984 doesn’t act alone. It works by looping through DNA’s three-dimensional structure to contact two genes that are critical for brain development: ETV5 and TRA2B.
Those DNA loops are strong in human brain tissue, but much weaker in chimpanzee, macaque, and mouse brains. In humans, the ETV5 protein also binds back to HAR1984, creating a feedback loop that keeps the enhancer active.
“This system helps fine-tune defining features of the human brain, including its size and the way it folds,” said Silver, senior study author and professor of cell biology, molecular genetics and microbiology and neurobiology at Duke.
The finding reinforces a growing idea in genetics: it’s not just which genes you have, but how DNA interact that shape who we are.
“The novelty here isn’t just the approach,” Silver said of the method first used in by her lab in 2025. “It’s the discovery that this enhancer actively influences how DNA interacts with its target genes to impact development.”
Why organoids aren’t enough
While brain organoids offer a powerful window into early human development, they have limitations.
Organoids can model many key cell types and processes, Silver said, but they still lack important features such as the blood vessel network that supply tissues with oxygen and nutrients and the full range of normal physiology.
That’s why the team paired organoid experiments with mouse studies to confirm that genetic effects hold up in living organisms.
What it means for evolution and disease
The study offers one of the clearest molecular explanations yet of how small, genetic changes – occurring only in humans – could have helped shape our uniquely capable brains.
It also highlights the importance of regulatory DNA, not just genes, in the story of human evolution.
“These findings point to specific DNA elements and cellular processes that influence human-specific brain development,” said Mosti. “Our work also helps identify other regions likely involved in human traits, including some linked to disease.”
Some HARs have already been statistically associated with neurodevelopmental conditions such as autism, raising the possibility that understanding these regions could one day inform medical research.
Silver and her team plan to study other HARs with similar patterns, including those in communication with each other. With more than 3,000 known HARs scattered across the genome, scientists believe HAR1984 is likely just one piece of a much larger puzzle.
“Understanding these enhancer regions helps us trace how the human brain became what it is today,” Silver said.
Support for the study was provided by the National Institutes of Health and a Triangle Center for Evolutionary Medicine award.
Additional Authors: Jing Liu, Katie Lam, Noah R. Dillon, Samantha Skavicus, Victoria A. Kapps, Ketrin Gjoni, Chia-Fang Lee, Emily N. Glidewell, Nicholas S. Heaton, and Katherine S. Polland.